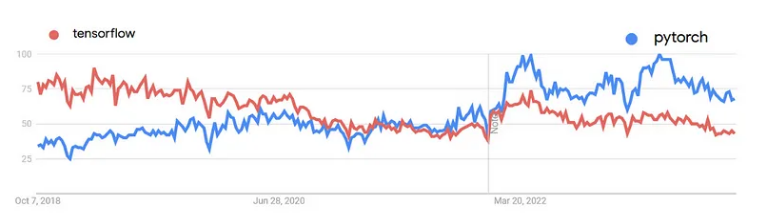
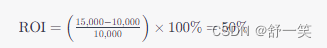第六节,我们使用结核病基因数据,做了一个数据预处理的实操案例。例子中结核类型,包括结核,潜隐进展,对照和潜隐,四个类别。第七节延续上个数据,进行了差异分析。 第八节对差异基因进行富集分析。本节进行WGCNA分析。
WGCNA分析 分段代码(附运行效果图)请查看上节
运行后效果
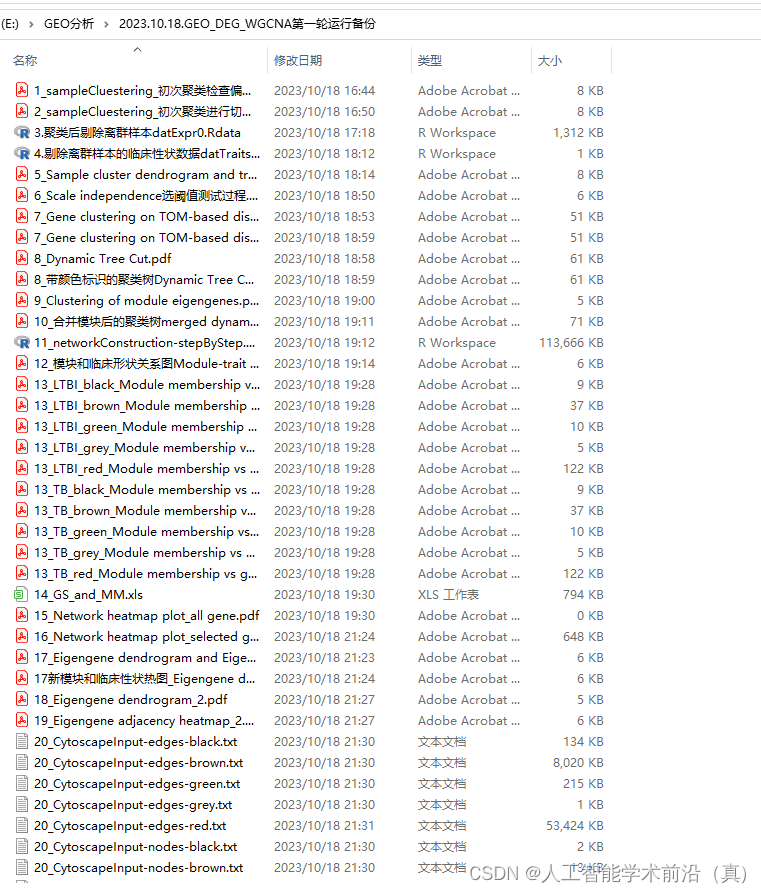
rm(list = ls()) ######清除环境数据#============================================================================
#======================================================================
#+========step0数据预处理和检查,已经做过step0==========================
#+========================================
#+=============================
"""
##############设置工作路径###################
workingDir = "C:/Users/Desktop/GSE152532"############工作路径,可以修改,可以设置为数据存放路径
setwd(workingDir)
getwd()################载入R包########################
library(WGCNA)
library(data.table)#############################导入数据##########################
# The following setting is important, do not omit.
options(stringsAsFactors = FALSE)
#Read in the female liver data set
fpkm = fread("Gene_expression.csv",header=T)##############数据文件名,根据实际修改,如果工作路径不是实际数据路径,需要添加正确的数据路径
# Take a quick look at what is in the data set
dim(fpkm)
names(fpkm)####################导入平台数据##########################
library(idmap2)
ids=get_soft_IDs('GPL10558')
head(ids)#####################将探针ID改为基因ID##########################
fpkm<-merge(fpkm,ids,by='ID')#merge()函数将dat1的探针id与芯片平台探针id相匹配,合并到dat1
library(limma)
fpkm<-avereps(fpkm[,-c(1,99)],ID=fpkm$symbol)#多个探针检测一个基因,合并一起,取其平均值
fpkm<-as.data.frame(fpkm)#将矩阵转换为表格
write.table(fpkm, file="FPKM_genesymbol.csv",row.names=T, col.names=T,quote=FALSE,sep=",")
###结束后查看文件,进行修改!!!# 加载自己的数据# load( "group_data_TB_LTBI.Rdata")load("exprSet_clean_mean_filter_log1.RData") #exprSet_cleanload( "dataset_TB_LTBI.Rdata")
exprSet_clean = dataset_TB_LTBI
gene_var <- apply(exprSet_clean, 1, var)##### 计算基因的方差
keep_genes <- gene_var >= quantile(gene_var, 0.75)##### 筛选方差较大的基因,选择方差前25%的基因
exprSet_clean <- exprSet_clean[keep_genes,]##### 保留筛选后的基因
dim(exprSet_clean)
save (exprSet_clean,file="方差前25per_TB_LTBI.Rdata")#######################基于方差筛选基因#################################
fpkm_var <- read.csv("FPKM_genesymbol.csv", header = TRUE, row.names = 1)##### 读入表达矩阵,矩阵的行是基因,列是样本
gene_var <- apply(fpkm_var, 1, var)##### 计算基因的方差
keep_genes <- gene_var >= quantile(gene_var, 0.75)##### 筛选方差较大的基因,选择方差前25%的基因
fpkm_var <- fpkm_var[keep_genes,]##### 保留筛选后的基因write.table(fpkm_var, file="FPKM_var.csv",row.names=T, col.names=T,quote=FALSE,sep=",")
###结束后查看文件,进行修改!!!##################重新载入数据########################
# The following setting is important, do not omit.
options(stringsAsFactors = FALSE)
#Read in the female liver data set
fpkm = fread("FPKM_var_filter.csv",header=T)##############数据文件名,根据实际修改,如果工作路径不是实际数据路径,需要添加正确的数据路径
# Take a quick look at what is in the data set
dim(fpkm)
names(fpkm)datExpr0 = as.data.frame(t(fpkm[,-1]))
names(datExpr0) = fpkm$ID;##########如果第一行是ID命名,就写成fpkm$ID
rownames(datExpr0) = names(fpkm[,-1])##################check missing value and filter ####################
load("方差前25per_TB_LTBI.Rdata")
datExpr0 = exprSet_clean##check missing value
library(WGCNA)
gsg = goodSamplesGenes(datExpr0, verbose = 3)
gsg$allOKif (!gsg$allOK)
{# Optionally, print the gene and sample names that were removed:if (sum(!gsg$goodGenes)>0)printFlush(paste("Removing genes:", paste(names(datExpr0)[!gsg$goodGenes], collapse = ", ")))if (sum(!gsg$goodSamples)>0)printFlush(paste("Removing samples:", paste(rownames(datExpr0)[!gsg$goodSamples], collapse = ", ")))# Remove the offending genes and samples from the data:datExpr0 = datExpr0[gsg$goodSamples, gsg$goodGenes]
}##filter
#meanFPKM=0.5 ####过滤标准,可以修改
#n=nrow(datExpr0)
#datExpr0[n+1,]=apply(datExpr0[c(1:nrow(datExpr0)),],2,mean)
#datExpr0=datExpr0[1:n,datExpr0[n+1,] > meanFPKM] # for meanFpkm in row n+1 and it must be above what you set--select meanFpkm>opt$meanFpkm(by rp)filtered_fpkm=t(datExpr0) #行 样本 列 基因
filtered_fpkm=data.frame(rownames(filtered_fpkm),filtered_fpkm)
names(filtered_fpkm)[1]="sample"
write.table(filtered_fpkm, file="FPKM_filter.csv",row.names=F, col.names=T,quote=FALSE,sep="\t")"""#&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&
#+&&&&&&&&&&&&&&&&&&加载数据&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
load('DEG_TB_LTBI_step13.Rdata') # DEG,res,all_diff,limma_clean_res,dataset_TB_LTBI_DEG,
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
#+&&&&&&&&&&&&&&&&&&加载数据&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&
#&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&library(WGCNA)
#读取目录名称,方便复制粘贴
dir()#============================================================================
#======================================================================
#+========step1样品聚类step1=================================
#+========================================
#+=============================################################样品聚类####################
#这里行是样品名,列为基因名,做转置处理
datExpr = t(dataset_TB_LTBI_DEG)
#初次聚类
sampleTree = hclust(dist(datExpr), method = "average")
# Plot the sample tree: Open a graphic output window of size 20 by 15 inches
# The user should change the dimensions if the window is too large or too small.
#设置绘图窗口
sizeGrWindow(12,9)
pdf(file='1_sampleCluestering_初次聚类检查偏离样本.pdf',width = 12,height = 9)
par(cex=0.6)
par(mar=c(0,4,2,0))
plot(sampleTree, main = "Sample clustering to detect outliers", sub="", xlab="", cex.lab = 1.5,cex.axis = 1.5, cex.main = 2)dev.off()#============================================================================
#======================================================================
#+========step2切割离群样本=================================
#+========================================
#+=============================pdf(file='2_sampleCluestering_初次聚类进行切割删除样本.pdf',width = 12,height = 9)
par(cex=0.6)
par(mar=c(0,4,2,0))
plot(sampleTree, main = "Sample clustering to detect outliers ", sub="", xlab="", cex.lab = 1.5,cex.axis = 1.5, cex.main = 2)### 测试画线,可以多次尝试
##############剪切高度问题,这个根据实际设置后可用
abline(h = 87, col = "red")##剪切高度不确定,故无红线
dev.off()### Determine cluster under the line
clust = cutreeStatic(sampleTree, cutHeight = 87, minSize = 10)
table(clust)
#clust
#0 1 2
#5 57 40
#由于本人案例,一刀切出三段,需要保留两段,用了’或‘逻辑运算符号
### 需要保留哪个,就传如要保留clust编号
keepSamples = (clust==1|clust==2)
#剔除离群样本
datExpr0 = datExpr[keepSamples, ]
#观察新表达矩阵
dim(datExpr0) #[1] 97 2813#&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&
#+&&&&&&&&&&&&&&&&&&数据保存&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
save(datExpr0,file='3.聚类后剔除离群样本datExpr0.Rdata')#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
#+&&&&&&&&&&&&&&&&&&数据保存&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&
#&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&load('datExpr0_cluster_filter.Rdata')#============================================================================
#======================================================================
#+========step3临床性状数据整理,与新表达矩阵保持一致=================================
#+========================================
#+=============================#加载自己的临床性状数据
load('design_TB_LTBI.Rdata')
traitData=designdim(traitData)# Form a data frame analogous to expression data that will hold the clinical traits.
fpkmSamples = rownames(datExpr0)
traitSamples =rownames(traitData)
#匹配样本名称,性状数据与表达数据保证一致
traitRows = match(fpkmSamples, traitSamples)
datTraits = traitData[traitRows,]
rownames(datTraits)
collectGarbage()#&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&
#+&&&&&&&&&&&&&&&&&&数据保存&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
save(datTraits,file='4.剔除离群样本的临床性状数据datTraits.Rdata')#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
#+&&&&&&&&&&&&&&&&&&数据保存&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&
#&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&#============================================================================
#======================================================================
#+========step4 增加临床性状数据后再次聚类=======================
#+========================================
#+=============================
# Re-cluster samples
sampleTree2 = hclust(dist(datExpr0), method = "average")
# Convert traits to a color representation: white means low, red means high, grey means missing entry
traitColors = numbers2colors(datTraits, signed = FALSE)
# Plot the sample dendrogram and the colors underneath.#sizeGrWindow(20,20)pdf(file="5_Sample cluster dendrogram and trait heatmap.pdf",width=12,height=12)
plotDendroAndColors(sampleTree2, traitColors,groupLabels = names(datTraits),main = "Sample dendrogram and trait heatmap")#Error in .plotOrderedColorSubplot(order = order, colors = colors, rowLabels = rowLabels, :
# Length of colors vector not compatible with number of objects in 'order'.dev.off()#============================================================================
#======================================================================
#+========step5 构建WGCNA网络=======================
#+========================================
#+=============================# Allow multi-threading within WGCNA. At present this call is necessary.
# Any error here may be ignored but you may want to update WGCNA if you see one.
# Caution: skip this line if you run RStudio or other third-party R environments.
# See note above.
#检查环境,能开几个线程
enableWGCNAThreads()# Choose a set of soft-thresholding powers
#设置阈值范围,WGCNA是无标度网络(scale free),节点连结数服从幂次定律分布。(连接数越多核心节点越少)
powers = c(1:15)# Call the network topology analysis function
#网络拓扑分析
sft = pickSoftThreshold(datExpr0, powerVector = powers, verbose = 5)# Plot the results:
sizeGrWindow(15, 9)
pdf(file="6_Scale independence选阈值测试过程.pdf",width=9,height=5)
#pdf(file="Rplot03.pdf",width=9,height=5)
par(mfrow = c(1,2))
cex1 = 0.9
# Scale-free topology fit index as a function of the soft-thresholding power
#无标度拓扑拟合指标作为软阈值能力的函数,根据下图结果,挑选合适阈值
plot(sft$fitIndices[,1], -sign(sft$fitIndices[,3])*sft$fitIndices[,2],xlab="Soft Threshold (power)",ylab="Scale Free Topology Model Fit,signed R^2",type="n",main = paste("Scale independence"));
text(sft$fitIndices[,1], -sign(sft$fitIndices[,3])*sft$fitIndices[,2],labels=powers,cex=cex1,col="red");
# this line corresponds to using an R^2 cut-off of h
abline(h=0.90,col="red")
# Mean connectivity as a function of the soft-thresholding power
plot(sft$fitIndices[,1], sft$fitIndices[,5],xlab="Soft Threshold (power)",ylab="Mean Connectivity", type="n",main = paste("Mean connectivity"))
text(sft$fitIndices[,1], sft$fitIndices[,5], labels=powers, cex=cex1,col="red")
dev.off()######chose the softPower
#选择阈值
softPower =sft$powerEstimate
adjacency = adjacency(datExpr0, power = softPower)##### Turn adjacency into topological overlap
#将邻接转换为拓扑重叠
TOM = TOMsimilarity(adjacency);
dissTOM = 1-TOM# Call the hierarchical clustering function
#无标度网络阈值参数确定后,调用分层聚类函数
#基于TOM的不相似性基因聚类
geneTree = hclust(as.dist(dissTOM), method = "average");
# Plot the resulting clustering tree (dendrogram)#sizeGrWindow(12,9)
pdf(file="7_Gene clustering on TOM-based dissimilarity基因聚类图.pdf",width=12,height=9)
plot(geneTree, xlab="", sub="", main = "Gene clustering on TOM-based dissimilarity",labels = FALSE, hang = 0.04)
dev.off()#聚类模块,最小的基因数量
# We like large modules, so we set the minimum module size relatively high:
minModuleSize = 30# Module identification using dynamic tree cut:
#使用dynamic tree cut进行模块识别
dynamicMods = cutreeDynamic(dendro = geneTree, distM = dissTOM,deepSplit = 2, pamRespectsDendro = FALSE,minClusterSize = minModuleSize);
table(dynamicMods)# Convert numeric lables into colors
#给不同模块分配颜色
dynamicColors = labels2colors(dynamicMods)
table(dynamicColors)
# Plot the dendrogram and colors underneath
#sizeGrWindow(8,6)
pdf(file="8_带颜色标识的聚类树Dynamic Tree Cut.pdf",width=8,height=6)
plotDendroAndColors(geneTree, dynamicColors, "Dynamic Tree Cut",dendroLabels = FALSE, hang = 0.03,addGuide = TRUE, guideHang = 0.05,main = "Gene dendrogram and module colors")
dev.off()# Calculate eigengenes
MEList = moduleEigengenes(datExpr0, colors = dynamicColors)
MEs = MEList$eigengenes
# Calculate dissimilarity of module eigengenes
MEDiss = 1-cor(MEs);
# Cluster module eigengenes
METree = hclust(as.dist(MEDiss), method = "average")
# Plot the result
#sizeGrWindow(7, 6)
pdf(file="9_Clustering of module eigengenes.pdf",width=7,height=6)
plot(METree, main = "Clustering of module eigengenes",xlab = "", sub = "")
MEDissThres = 0.25######剪切高度可修改
# Plot the cut line into the dendrogram
abline(h=MEDissThres, col = "red")
dev.off()# Call an automatic merging function
#根据前面设置的剪切高度,对模块进行合并
merge = mergeCloseModules(datExpr0, dynamicColors, cutHeight = MEDissThres, verbose = 3)
# The merged module colors
mergedColors = merge$colors
# Eigengenes of the new merged modules:
mergedMEs = merge$newMEs#sizeGrWindow(12, 9)
pdf(file="10_合并模块后的聚类树merged dynamic.pdf", width = 9, height = 6)
plotDendroAndColors(geneTree, cbind(dynamicColors, mergedColors),c("Dynamic Tree Cut", "Merged dynamic"),dendroLabels = FALSE, hang = 0.03,addGuide = TRUE, guideHang = 0.05)
dev.off()# Rename to moduleColors
moduleColors = mergedColors
# Construct numerical labels corresponding to the colors
#构建相应颜色的数字标签
colorOrder = c("grey", standardColors(50))
moduleLabels = match(moduleColors, colorOrder)-1
MEs = mergedMEs# Save module colors and labels for use in subsequent parts
#&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&
#+&&&&&&&&&&&&&&&&&&数据保存&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
save(datExpr0,datTraits,MEs, TOM, dissTOM, moduleLabels, moduleColors, geneTree, sft, file = "11_networkConstruction-stepByStep.RData")
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
#+&&&&&&&&&&&&&&&&&&数据保存&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&
#&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&&load("11_networkConstruction-stepByStep.RData")#=====================================================================================
#===============================================================================
#+========step6 计算模块和临床性状相关系数(核心挑选色块)==============
#+========================================
#+=============================
##############################relate modules to external clinical triats# Define numbers of genes and samples
nGenes = ncol(datExpr0)
nSamples = nrow(datExpr0)#可以修改参数 p值pvalue 更换
moduleTraitCor = cor(MEs, datTraits, use = "p")
moduleTraitPvalue = corPvalueStudent(moduleTraitCor, nSamples)#sizeGrWindow(10,6)
pdf(file="12_模块和临床形状关系图Module-trait relationships.pdf",width=10,height=6)
# Will display correlations and their p-values
textMatrix = paste(signif(moduleTraitCor, 2), "\n(",signif(moduleTraitPvalue, 1), ")", sep = "")dim(textMatrix) = dim(moduleTraitCor)
par(mar = c(6, 8.5, 3, 3))# Display the correlation values within a heatmap plot #修改性状类型 data.frame
labeledHeatmap(Matrix = moduleTraitCor,xLabels = names(data.frame(datTraits)),yLabels = names(MEs),ySymbols = names(MEs),colorLabels = FALSE,colors = greenWhiteRed(50),textMatrix = textMatrix,setStdMargins = FALSE,cex.text = 0.5,zlim = c(-1,1),main = paste("Module-trait relationships"))
dev.off()#色块 red相关度 0.75#=====================================================================================
#===============================================================================
#+========step7 定义包含所有datTraits列的可变权重(MM and GS)==============
#+========================================
#+=============================#定义包含所有datTraits列的可变权重######## Define variable weight containing all column of datTraits###MM(gene Module Membership) and GS(gene Trait Significance)# names (colors) of the modules
modNames = substring(names(MEs), 3)geneModuleMembership = as.data.frame(cor(datExpr0, MEs, use = "p"))
MMPvalue = as.data.frame(corPvalueStudent(as.matrix(geneModuleMembership), nSamples))names(geneModuleMembership) = paste("MM", modNames, sep="")
names(MMPvalue) = paste("p.MM", modNames, sep="")#names of those trait
traitNames=names(data.frame(datTraits))
class(datTraits)geneTraitSignificance = as.data.frame(cor(datExpr0, datTraits, use = "p"))
GSPvalue = as.data.frame(corPvalueStudent(as.matrix(geneTraitSignificance), nSamples))names(geneTraitSignificance) = paste("GS.", traitNames, sep="")
names(GSPvalue) = paste("p.GS.", traitNames, sep="")####plot MM vs GS for each trait vs each module##########example:royalblue and CK
module="red"
column = match(module, modNames)
moduleGenes = moduleColors==moduletrait="TB"
traitColumn=match(trait,traitNames)sizeGrWindow(7, 7)#par(mfrow = c(1,1))
verboseScatterplot(abs(geneModuleMembership[moduleGenes, column]),
abs(geneTraitSignificance[moduleGenes, traitColumn]),
xlab = paste("Module Membership in", module, "module"),
ylab = paste("Gene significance for ",trait),
main = paste("Module membership vs. gene significance\n"),
cex.main = 1.2, cex.lab = 1.2, cex.axis = 1.2, col = module)
######for (trait in traitNames){traitColumn=match(trait,traitNames)for (module in modNames){column = match(module, modNames)moduleGenes = moduleColors==moduleif (nrow(geneModuleMembership[moduleGenes,]) > 1){####进行这部分计算必须每个模块内基因数量大于2,由于前面设置了最小数量是30,这里可以不做这个判断,但是grey有可能会出现1个gene,它会导致代码运行的时候中断,故设置这一步#sizeGrWindow(7, 7)pdf(file=paste("13_", trait, "_", module,"_Module membership vs gene significance.pdf",sep=""),width=7,height=7)par(mfrow = c(1,1))verboseScatterplot(abs(geneModuleMembership[moduleGenes, column]),abs(geneTraitSignificance[moduleGenes, traitColumn]),xlab = paste("Module Membership in", module, "module"),ylab = paste("Gene significance for ",trait),main = paste("Module membership vs. gene significance\n"),cex.main = 1.2, cex.lab = 1.2, cex.axis = 1.2, col = module)dev.off()}}
}#####
names(datExpr0)
probes = names(data.frame(datExpr0))#=====================================================================================
#===============================================================================
#+========step8 导出计算完毕的(MM and GS)==============
#+========================================
#+=============================
#################export GS and MM############### geneInfo0 = data.frame(probes= probes,moduleColor = moduleColors)for (Tra in 1:ncol(geneTraitSignificance))
{oldNames = names(geneInfo0)geneInfo0 = data.frame(geneInfo0, geneTraitSignificance[,Tra],GSPvalue[, Tra])names(geneInfo0) = c(oldNames,names(geneTraitSignificance)[Tra],names(GSPvalue)[Tra])
}for (mod in 1:ncol(geneModuleMembership))
{oldNames = names(geneInfo0)geneInfo0 = data.frame(geneInfo0, geneModuleMembership[,mod],MMPvalue[, mod])names(geneInfo0) = c(oldNames,names(geneModuleMembership)[mod],names(MMPvalue)[mod])
}
geneOrder =order(geneInfo0$moduleColor)
geneInfo = geneInfo0[geneOrder, ]write.table(geneInfo, file = "14_GS_and_MM.xls",sep="\t",row.names=F)#=====================================================================================
#===============================================================================
#+========step9 基因网络热图进行可视化(非常耗费内存资源)==============
#+========================================
#+=============================nGenes = ncol(datExpr0)
nSamples = nrow(datExpr0)# Transform dissTOM with a power to make moderately strong connections more visible in the heatmap
plotTOM = dissTOM^7
# Set diagonal to NA for a nicer plot
diag(plotTOM) = NA# Call the plot functionsizeGrWindow(9,9) #这个耗电脑内存
pdf(file="15_所有基因数量太多Network heatmap plot_all gene.pdf",width=9, height=9)
TOMplot(plotTOM, geneTree, moduleColors, main = "Network heatmap plot, all genes")
dev.off()nSelect = 400
# For reproducibility, we set the random seed
set.seed(10)
select = sample(nGenes, size = nSelect)
selectTOM = dissTOM[select, select]
# There's no simple way of restricting a clustering tree to a subset of genes, so we must re-cluster.
selectTree = hclust(as.dist(selectTOM), method = "average")
selectColors = moduleColors[select]# Open a graphical window
#sizeGrWindow(9,9)
# Taking the dissimilarity to a power, say 10, makes the plot more informative by effectively changing
# the color palette; setting the diagonal to NA also improves the clarity of the plot
plotDiss = selectTOM^7
diag(plotDiss) = NApdf(file="16_400个基因试试Network heatmap plot_selected genes.pdf",width=9, height=9)
TOMplot(plotDiss, selectTree, selectColors, main = "Network heatmap plot, selected genes")
dev.off()#=====================================================================================
#===============================================================================
#+========step10 新模块和临床性状热图 合并和拆分两个版本==============
#+========================================
#+=============================#sizeGrWindow(5,7.5)
pdf(file="17新模块和临床性状热图_Eigengene dendrogram and Eigengene adjacency heatmap.pdf", width=5, height=7.5)
par(cex = 0.9)
plotEigengeneNetworks(MEs, "", marDendro = c(0,4,1,2), marHeatmap = c(3,4,1,2), cex.lab = 0.8, xLabelsAngle= 90)
dev.off()#or devide into two parts
# Plot the dendrogram
#sizeGrWindow(6,6);
pdf(file="18_Eigengene dendrogram_2.pdf",width=6, height=6)
par(cex = 1.0)
plotEigengeneNetworks(MEs, "Eigengene dendrogram", marDendro = c(0,4,2,0), plotHeatmaps = FALSE)
dev.off()pdf(file="19_Eigengene adjacency heatmap_2.pdf",width=6, height=6)
# Plot the heatmap matrix (note: this plot will overwrite the dendrogram plot)
par(cex = 1.0)
plotEigengeneNetworks(MEs, "Eigengene adjacency heatmap", marHeatmap = c(3,4,2,2), plotDendrograms = FALSE, xLabelsAngle = 90)
dev.off()###########################Exporting to Cytoscape all one by one ###########################=====================================================================================
#===============================================================================
#+========step11 导出每个模块的边和节点关系(Cytoscape 绘图所需)==============
#+========================================
#+=============================# Select each module
'''
Error in exportNetworkToCytoscape(modTOM, edgeFile = paste("CytoscapeInput-edges-", : Cannot determine node names: nodeNames is NULL and adjMat has no dimnames.datExpr0 格式需要dataframe
'''
modules =module #module="red"
for (mod in 1:nrow(table(moduleColors)))
{modules = names(table(moduleColors))[mod]# Select module probesprobes = names(data.frame(datExpr0)) # inModule = (moduleColors == modules)modProbes = probes[inModule]modGenes = modProbes# Select the corresponding Topological OverlapmodTOM = TOM[inModule, inModule]dimnames(modTOM) = list(modProbes, modProbes)# Export the network into edge and node list files Cytoscape can readcyt = exportNetworkToCytoscape(modTOM,edgeFile = paste("20_CytoscapeInput-edges-", modules , ".txt", sep=""),nodeFile = paste("20_CytoscapeInput-nodes-", modules, ".txt", sep=""),weighted = TRUE,threshold = 0.02,nodeNames = modProbes,altNodeNames = modGenes,nodeAttr = moduleColors[inModule])
}WGCNA关系网络的构建完毕,绘图找核心基因,Cytoscape 到底怎么玩?